Yeast terminology, part 2: bacteria
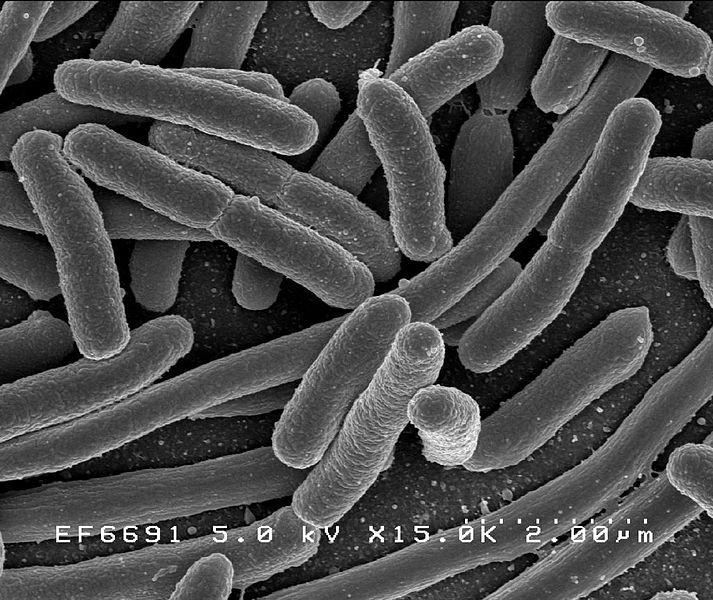
Escherichia coli bacteria, from Wikimedia Commons |
We continue the series on the family tree of yeast with a post on bacteria. As I explained in the first post, bacteria are very different from yeast, but they are still important in beer. (Yes, I should have thought of a better title while I was still doing part 1. Ah well.)
People usually don't think of bacteria at all, or if they do, they're considered the bringers of disease, tooth decay, and rot. But really, bacteria are the true rulers of this planet, and we are just guests along for the ride. Bacteria turned earth into a planet with an oxygen athmosphere, they run many of the essential chemical feedback loops that keep the ecosystem working, and they exist absolutely everywhere that you can find heat, nutrients, and water.
This very much includes your own body, which contains more bacterial cells than human cells, so strictly speaking you are a symbiosis between eukaryotes and bacteria. If the flora of bacteria in your gut gets upset you may have to spend years going in and out of hospitals. Bacteria are by no means a sideshow in life on earth, and so of course they have a role to play in beer as well.
Bacteria are often divided into gram-positive and gram-negative species. These differ in the structure of their cell walls, which means that they can be told apart with a simple colour test, called gram staining. Essentially, the bacteria are given purple dye, which they absorb, and are then exposed to alcohol and red dye. This makes the gram-negative bacteria lose the purple dye through their thinner cell walls, and absorb the red dye. The gram-positive retain the purple dye, and appear purple under the microscope, whereas the gram-negative lose it and instead appear red.
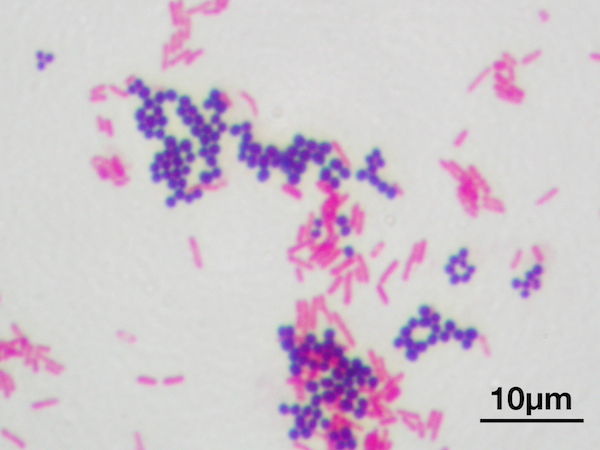
Gram staining. Violet bacteria are gram-positive, pink are gram-negative. Y tambe, CC BY SA |
It's a simple test, but it happens to correspond to a deep taxonomic divide between different kinds of bacteria, and so is a very useful tool for microbiologists. In 1937, the Irish microbiologist J. L. Shimwell discovered that hops prevent gram-positive bacteria from growing, but not the gram-negative, so this divide is deeply relevant for beer.
Some relevant gram-negative bacteria are Enterobacter, Klebsiella, Shigella, and Hafnia, all of which occur in lambic fermentation. These can also spoil ordinary beer, but they are not the chief culprits. Note that all these single names are not species but genera (plural of genus); species have two names, like Hafnia alvei.
The gram-negative bacteria contain one larger grouping that is of interest, and that is the family of Acetobacteraceae, or acetic acid bacteria. These bacteria are rod-shaped and produce acetic acid, which is the main acid in vinegar, and has a sharply sour flavour and a strong smell. They are generally not wanted in beer, but do occur in lambic fermentations and are sometimes used to sour Flemish red ale. The most important ones are Acetobacter and Gluconobacter.
Some important gram-positive genera of bacteria are Staphylococcus, Streptococcus, Clostridium, and Listeria, which contain many species that are harmful to human beings. Just as well that the hops prevent them.
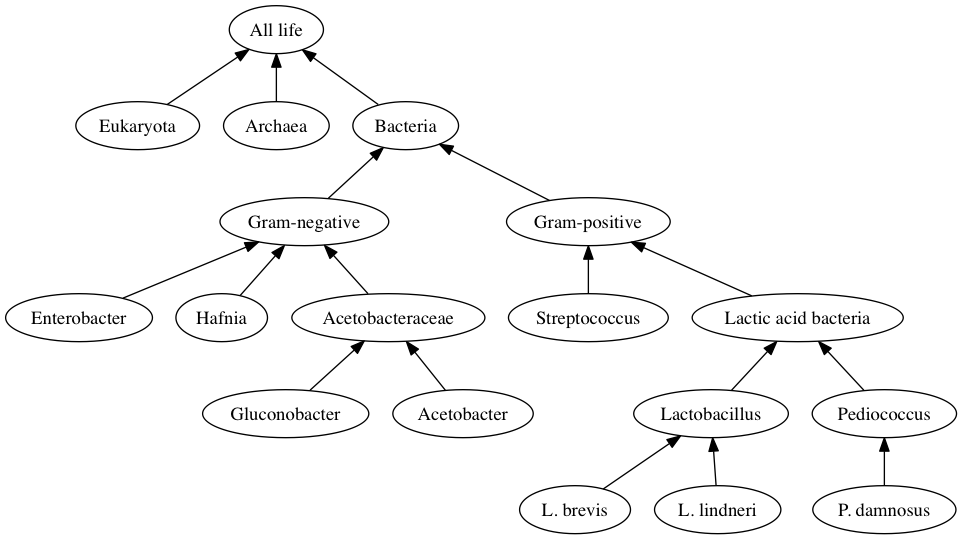
Taxonomic tree of bacteria |
Weirdly, the bacteria that are well known as either spoilers of beer or which are sometimes added deliberately are nearly all gram-positive. The hops ought to kill them, but in fact they don't, because some strains of gram-positive bacteria have developed hop resistance. Note that I really do mean "strains" here, because within most of the gram-positive species only some of the strains have hop resistance.
Nearly all the gram-positive bacteria that appear in beer belong to a group called the lactic acid bacteria. As the name suggests, they produce lactic acid, which is a gentler, cleaner-tasting acid than acetic acid. The sour German wheat beers, like Berliner weisse and gose are soured with this type of bacteria, and of course they also occur in lambic fermentations. They are also the most common beer spoilers.
The best-known genus is Lactobacillus, which contains an enormous number of species. They live in humans, and are widely used in food, such as yogurt, cheese, etc. The most important species here is Lactobacillus brevis, which according to some analyses is responsible for up to half of all cases of infected beer. Lactobacillus lindneri is another common offender, and it's unusual in that all L. lindneri strains tested so far are hop-resistant.
Another important genus in this group is Pediococcus, which tends to produce diacetyl and can form slimy ropes. The most common beer spoiler in this genus is Pediococcus damnosus, which also dominates the late stage of lambic fermentation. One strain is actually sold by White Labs for use in beer.
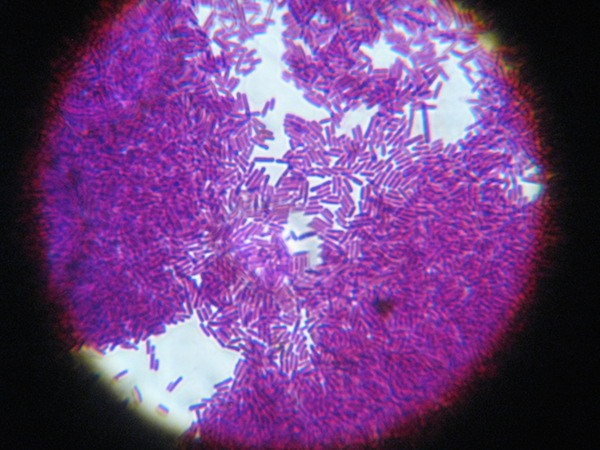
Lactobacillus, by Shoko Muraguchi, BY NC ND |
Two more blog posts to go in this series. One will look at the Saccharomyces genus, and another at exactly how hops prevent bacterial infections.
Similar posts
How hops prevent infection
The increasingly inaccurately named series on yeast terminology continues with a post diving into how, exactly, hops prevent bacteria from infecting beer
Read | 2015-09-18 14:43
What is it that ferments lambic?
As everyone knows, lambic is fermented by "wild yeast and bacteria"
Read | 2015-02-20 11:00
Yeast terminology, part 1
I was asked to explain the family trees of yeast and since not much has been written on this I figured I'd give it a go
Read | 2015-08-27 18:54
Comments
Krister G - 2015-09-05 19:36:08
This is a great introduction, very easy to understand with even a minimal knowledge of microbiology! I will definetly read the rest of these posts, and follow the blog in the future.
I would like to know one thing though; is there a formula used to determine how much hops you need to add in order to minimise the risk of infection? / -Also to make sure you don't kill off your added bacteria...
Anyway, great to see someone covering this part of brewing in a more detailed fashion!
Are Gulbrandsen - 2015-09-17 18:06:35
Any news about the Hornindal kveik bacteria?
Lars Marius - 2015-09-18 03:44:42
@Krister: The next blog post goes into this, but basically the bitterness of the beer and the bacteria protection are one and the same. It's both done by the alpha acid. The bacteria you deliberately add to your beer are hop resistant. More about that, too, in the next blog post.
@Are: We don't know what they are yet, but the guys at NTNU say they are going to try to identify them. If they succeed I will be posting about it both here and on Twitter.