A family tree for brewer's yeast

Yeast starter, Aizpute, Latvia |
I wrote a series of blog posts on the family tree of yeast, starting at the very top and continuing all the way down to the family of yeast species. The story doesn't end there, however, because within the species of Saccharomyces cerevisiae (ale yeast) the different strains form a family tree of their own. However, when I wrote those blog posts in 2015 the shape of this tree was poorly understood.
That changed with the publication of a landmark paper in Cell in 2016[1], where no less than 450 yeast strains were assembled into one big family tree. That really did give us an outline of the family tree, and the results were fairly startling, so I want to try to share them.
What and how
The researchers started by collecting 157 strains of yeast. All of these belonged to the same species, Saccharomyces cerevisiae, which is the yeast species used to ferment most beers, but not lager. There is a family tree of strains because even though they belong to the same species they don't all have the same genes. Think about human beings, Homo sapiens. Hair colour, hair shape, skin colour, and so on, all differ because of genetic differences. Some people cannot smell acetaldehyde because they lack the necessary gene, while others can.

Fermenting beer, Pihtla, Saaremaa, Estonia |
It's the same with yeast. All strains of ale yeast are broadly similar, but not the same. So basically, two yeast cells belong to the same strain if they are genetically identical. Or nearly identical.
The researchers sequenced the genes of all these yeast strains. That is, they decoded their genes, so they could read them as sequences of one of the four molecules that code the genes, something like this: AACTGGCA... They sequenced the entire genome, which is about 6300 genes, but only used about a third of it, 2020 genes. They deliberately left out the non-coding part of the genome, because (I'm guessing) they considered that irrelevant mutations in the non-functional part would be noise which could disturb the results.
They then used the genomes to build a family tree by looking at which strains had the most similar set of genes to which others. The ones that were the most similar to each other were assumed to be closely related, and the more different ones to be more distantly related. Using this approach they were able to have a computer algorithm build a family tree. Then they extended the tree with 293 more strains, which they had less genetic data on, but still enough to build a reasonable tree.
The actual tree
So, let's turn to the actual tree. Note that the researchers only put in the genes, without telling the computer what sort of yeast they came from. The collection contained not just beer yeast, but also wine yeast, Asian rice wine yeasts, wild yeasts, and even yeast collected from infected human beings. The output was:
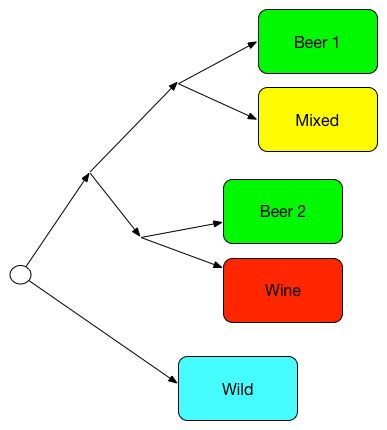
Simplified family tree, redrawn after Gallone et al[1] |
Two big groups are called "Beer 1" and "Beer 2", where "Beer 2" is more closely related to the "Wine" group than to the first group, and the first group is more closely related to the "Mixed" group. The "Wild" group is the furthest away from all the others.
This suggests that most brewing yeasts fall into one of two different groups. (There were some beer yeasts that didn't land in either "Beer 1" or "Beer 2".) Unfortunately, the paper does not identify the individual strains, and it says very little about the differences between the two groups, so this is less useful than it might sound, but it's still fascinating. Let's hope more details on the two groups will be forthcoming with time.
From the diagrams in the paper we can deduce some differences between the two groups:
- The Beer 2 yeasts tend to have higher alcohol tolerance.
- The Beer 1 yeasts are better at fermenting maltotriose.
- The Beer 2 yeasts handle drying better.
However, it gets more intriguing that this, because variation within "Beer 1" is so small that the researchers concluded that the whole group must derive from a single ancestor that lived 3-4 centuries ago. The result for "Beer 2" is much the same. That's startling, because it implies that nearly all of the yeasts brewers use today descend from just two original strains of yeast back in the 1600s.
That's really bizarre. Think about how many different ale breweries there have been over the last 3-4 centuries, all over Ireland, the UK, France, Belgium, the Netherlands, Germany, Scandinavia, etc etc. All of these have used one or more yeast strains that they got from some other brewer when starting up (except the very few breweries using spontaneous fermentation). Seemingly they have all used descendants of the same two strains. How on earth could that happen?
I'll try to return to that in a later blog post. Meanwhile, let's look at another fascinating insight.
Domestication
The researchers also looked at which genes actually make the differences between these groups. Let's start with the "Wild" group. These are yeasts that live in nature. This is where all the yeast strains started, but some gradually moved into breweries and by now descend from yeasts that lived there for centuries, without ever venturing back into the wild. So how did the genes of the "domesticated" yeasts change?
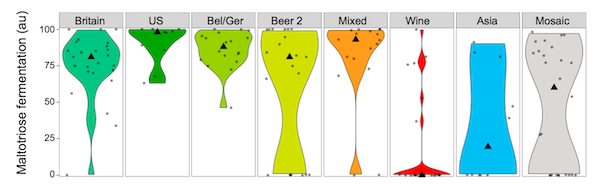
Maltotriose utilization in the various yeast groups, from Gallone et al[1], figure 3E, p1403. |
The "Wine" group shows the smallest changes from the wild. Basically, they lost their ability to ferment maltotriose (a sugar from malts, which doesn't exist in wine), and they became more tolerant to the kinds of stresses yeast experience when fermenting wine must. Other than that they don't seem to have changed that much. They're still good at surviving in nature, for example. So, basically, the wine yeasts are domesticated, but less so than the beer yeasts.
This makes sense, because wine is made only once per year (when the grapes are ripe), and historically the yeast would then spend the rest of the year in the wild, before fermenting next year's vintage. So wine has been through fewer generations of fermenting wine, and even spent at least some time in the wild in between.
The wine yeasts also turn out to have high tolerance for sulfites and copper, both of which are used in wine production. The other yeast groups have visibly less tolerance for copper and sulfites.
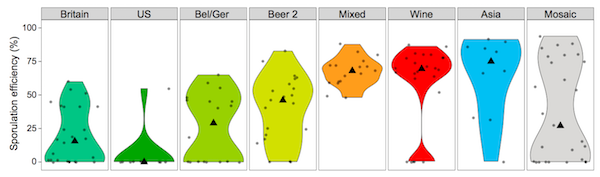
Sporulation efficiency in the various yeast groups, from Gallone et al[1], figure 4A, p1405. |
The beer yeasts show much bigger changes. They have become better at fermenting maltotriose, they have lost the genes for off flavours, they survive poorly in nature (see image above), and parts of the genome have decayed. This is what you'd expect from an organism that has spent thousands of generations in an environment that is far less harsh than nature, where it is taken care of, fed, and where humans actively select the strains they like best. Parts of the genome decay simply because they are no longer needed. For example, the genes that enable these yeasts to survive drying are crucial in nature and next to useless in a brewery.
The flavour change is interesting. Two genes, PAD1 and FDC1, control the production of 4-vinyl guaiacol (4-VG), a phenolic off-flavour. The wild yeasts generally have these, but in the Beer 1 and 2 groups these genes have usually been destroyed by nonsense mutations. Analysis of the tree suggests that these mutations happened quite early in the history of these yeasts. So very likely humans actively selected, kept, and perhaps shared the yeasts that didn't make these flavours, and threw away the ones that did.
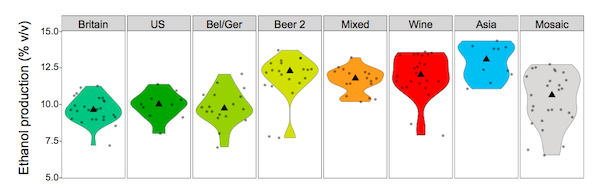
Alcohol tolerance in the various yeast groups, from Gallone et al[1], figure 3A, p1403. |
The way the yeasts have been shaped by how humans use them is shown very clearly in the image above. The wine and "Asia" yeasts (sake, rice wine, etc) have the highest alcohol tolerance, which is entirely logical, since these are the strongest drinks.
Note that what you are looking at here is actually evolution in practice. A combination of mutations, natural variation, natural selection, and some human selection have combined to change living organisms to make them better adapted to their environment, whether that is a brewery, winery, or a sake brewery. The researchers estimate that the changes to the beer yeasts have taken about 75,000 generations, so we're looking a time span that if translated to human generations would be about 1,500,000 years.
History
The paper has even more to offer than this, because the researchers tried to estimate how long ago the last common ancestors in each of the groups Beer 1 and Beer 2 lived. Based on the rate of mutation and some assumptions about the number of yeast generations per year, they estimate that Beer 1 started diverging some time between 1573-1604 CE, and Beer 2 some time 1645-1671 CE. (Note that Beer 1 and Beer 2 separated far earlier than that.) That's astonishingly recent, for reasons that we will return to.
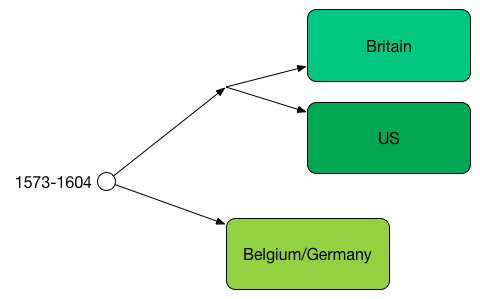
Family tree within "Beer 1", redrawn from Gallone et al[1], figure 1A, p1399. |
The researchers then looked at where the yeasts came from, and how that fit into the tree. In Beer 1, the deepest split is between Belgian and German yeast on one side, and British and American on the other side. The Belgian and the German yeasts are mixed together, but there is an American subgroup, distinct from the British.
This would seem to imply that a very large proportion of the brewing yeasts in these countries all descend from a single strain. So breweries somewhere in Europe must have, somehow, gotten hold of a strain that was then shared throughout four different countries, and that was also taken across the Atlantic to the US. You really have to wonder how and why that happened.
I used to think that the yeasts sold as "American ale yeast" were really European yeasts that crossed the Atlantic in the last few decades and that considering them American was more of a marketing ploy than strict historical fact. The results in this paper, however, indicate that there really is a separate lineage of yeasts that have developed in America over a substantial period of time.
Problematic history
The authors also write some things I consider unfortunate, because in my opinion they further misunderstandings about the history of yeast. For example, they write that "because early brewers, winemakers, and bakers were unaware of the existence of yeast, there is no record of how yeasts made their way into these processes." This is not true at all. We have accounts from the 16th and 17th centuries telling us exactly this. Also, people might not have known what yeast was in precise biological terms, but they knew it existed, they had words for it, equipment for using it, and they knew how to use it.
To this the researchers respond:
Yes, 16th century brewers kind of knew what yeast is, in the sense that they learned to reclusive the sediment (mostly yeast) and had words to describe this. But, they did likely not realize that this material consisted of living organisms, microbes, and that they were selecting and domesticating these organisms (in contrast to pets and livestock). So, that is the point we wanted to make in the paper.
I agree that brewers at this time probably were not aware that yeast was a living organism, and they definitely were not breeding it deliberately the way they did with livestock. The interesting thing (which is the point the authors were really making) is that brewers effectively did breed these yeasts without knowing it.
Later, they write that the start date for Beer 1 (1573-1604) suggests "that domestication started around this time. Interestingly, this coincides with the gradual switch from home-centered beer brewing where every family produced their own beer, to more professional large-scale brewing, first in pubs and monasteries and later also in breweries."

Site of 17th century brewery, Leuven, Belgium |
However, large-scale brewing began much earlier, as William Unger documents[2]. In the 13th century beer was a major export article and income from taxation on it was very important for nation states and city states. Already in the 15th century commercial brewers were using copper kettles of 4000-5000 liters. That is definitely professional large-scale brewing, well over a century before 1573.
The authors responded to this, too, as follows:
We would like to clarify that with large-scale brewing and industrial breweries, we are not referring only to the appearance of relatively big breweries (probably already present before 1500) but to the industrialization of the brewing industry, including intensified trade across long distances. I think it's relevant to mention this passage in the Comprehensive history of beer brewing, 2009 by FG Meussdoerffer, p18-19:
"By the sixteenth century the infrastructure in Central Europe had improved so far that traveling and transport became possible at speeds not encountered since he times of the Roman legions. Thus, overland transport became feasible, even for commodities like beer, although at low volumes and high costs."
I think the authors make a reasonable point here, and in the end it comes down to how big a brewery needs to be to be considered big, which is obviously subjective. However, I believe domestication of yeast very likely began so long before this time frame that the discussion is moot. The evidence is a bit involved, so that will have to be a subject for a different blog post.
And kveik?
So how does kveik and the other farmhouse yeasts, such as the Lithuanian and the Chuvashian ones, fit into this? Is kveik "Beer 3"? Right now, nobody has any idea at all. However, since these very likely come out of a completely different historical background, chances are that fitting them into the tree could tell us a lot more about the history of yeast. So far, nobody's actually looking at that. Unfortunately. A couple of groups of researchers are hoping to gather the necessary funds to find out. Let's hope they succeed.
References
[1] Domestication and Divergence of Saccharomyces cerevisiae Beer Yeasts, Gallone et al., 2016, Cell 166, 1397–1410 September 8, 2016 http://dx.doi.org/10.1016/j.cell.2016.08.020
[2] A History of Brewing in Holland 900-1900, Richard W. Unger, Brill, 2001. ISBN 9004120378, p15-20.
Acknowledgements
Many thanks to Anders Christensen and Richard Preiss for very good discussions of this paper that helped me understand it better.
Also very warm thanks to Kevin Verstrepen, Brigida Gallone, Jan Steensels, and Steven Maere (authors of the paper) for taking the time to reply so carefully to the first draft of this blog post.
Similar posts
A family tree for kveik
In 2016 I was contacted by Canadian researcher Richard Preiss
Read | 2017-10-06 10:02
Where kveik comes from
I've written before about the kveik research paper by Preiss, Tyrawa, and van der Merwe
Read | 2018-09-12 16:27
Studying Farmhouse Yeast Genomes
This is the third blog post on the new Verstrepen lab paper on farmhouse yeast
Read | 2026-05-03 13:59
Comments
Dan Pixley - 2017-08-08 23:31:13
I have a joke for you, "A historian and a microbiologist walk into a bar..."
Lars Marius Garshol - 2017-08-09 05:39:39
Hmmmmmm. I don't think I get it. :-)
Tyson - 2017-08-10 05:40:34
Should publish some links for the researchers who need funding for the next phase. I'm sure the beer community could do a lot for making it happen. I'd post it up in my small store and put at least something in. I already have the beer periodic table and a few other discussion building items up.
Ulrich Bähr - 2017-08-11 09:41:50
Excellent paper! I already knew the orginal paper - but now I understand it :-) Thank you.
A short comment on the improved logistics infrastructure in Central Europe: The celts apparently layed out the roads we now know as Roman Roads, the Romans paved it (which certainly sped up travelling). In the 8th century notable quantities of wine were transported from Italy over the Alps to the Danube. Improved transportation facilities cannot really explain the centralization of beer production in the Late Middle Ages.
Lars Marius Garshol - 2017-08-11 10:12:14
@Tyson: It doesn't seem like the researchers themselves want to work this way. Unfortunately. Otherwise I'd agree that this seems like a good way to fund it.
@Ulrich: I'm tempted to agree. Wine was exported from the Mediterranean to Scandinavia long before the birth of Christ. So it's difficult to see that anything changed very much around 1500. And even long after then most of the export of beer appears to have gone by ship.
qq - 2017-08-14 11:27:13
The kind of DNA you use for this kind of study depends on the timescale you are studying. For instance if you are studying recent changes in DNA, such as changes that have happened since humans became a species because eg police want to identify individual people ("DNA fingerprinting") then you want DNA that changes fast, because it has little selection pressure, so you use "junk" DNA. If you're looking at the deep relationships between eg vertebrates and echinoderms over 100's of millions of years then you want DNA that changes very slowly because it codes really important genes where mutations are usually fatal, things like the basic mechanism for making proteins. For this kind of study you want the genes in between, that have some selection pressure but are not fatal if they change - things like skin colour or lactose tolerance in humans.
I wouldn't take the dates too literally - all the geneticists can "see" are generations, and the dates depend heavily on what assumptions you make about how many generations the main lineage experienced per year. They seem to assume about 200 generations per year - but the main lineage will see far fewer than that if you're doing seasonal brewing (eg Märzen) or storing kveik on a ring, and in turn that will push the split further back in time.
It is intriguing to see so few families of yeast though, when yeast is a ubiquitous thing - you can always find some kind of yeast for a wild fermentation. That implies the spread of a certain beer culture that was so obviously "better" than earlier "farm" beers. The fact that this seems to have happened in the late Middle Ages suggests that the spread of domestic yeast seems to have happened alongside the big change in beer culture at that time - hops. It would be interesting to try and map where the new yeast appeared, I suspect domestic yeast and hops are very closely related in time and space.
PS Lars - I owe you thanks, we were talking at the Great British Beer Festival about Lithuanian beer, and reading your stuff meant that I didn't sound a complete idiot! :-)
Lars Marius Garshol - 2017-08-18 15:24:48
@qq: Thank you for giving more detail on which DNA to choose. That makes a lot of sense.
I haven't taken the dates too literally, but I didn't want to start discussing whether I agreed or not with them in this post, because it's already more than long enough. But, yes, 50 brews per year is a highly debatable assumption.
Your point about hops is interesting, and matches the distribution somewhat. That's worth pondering further.
Glad to hear you spoke about Lithuanian beer! More people need to do that. And I'm glad my writings could be of use.
qq - 2017-12-06 00:20:53
Suregork's been able to identify a couple of the coded strains to White Lab yeast by slogging through huge amounts of DNA data, I've then come in and made some informed guesses about which other White Labs strains are which - they're mostly in a big block in Beer44-86.
http://beer.suregork.com/?p=3907#comment-600297
It's not confirmed yet - unless the Leuven lab wants to say something or someone does some PCRs using the genome sequences as a guide - but it all seems to fit. Which means we have genomes and a bunch of other data about most of the White Labs ale strains! :-)))))
Mads Leonard Holvik - 2022-09-20 18:10:38
Thanks for great articles! My instinct about people who brewed beer let's say in the 1600's, and who saw the fermentation process going on in the fermentation vessels and who used the yeast again at next fermentation, certainly knew that this is something alive. I don't understand why the scientists would assume that people had no idea that the beer yeast is alive.
siddhi thaware - 2024-04-18 14:22:50
360iResearch recently published a market research report titled "Brewer’s Spent Yeast Market by Type (Dry, Liquid), Category (Primary Spent Yeast, Secondary Spent Yeast), Application - Global Forecast 2024-2030". This comprehensive report provides a detailed analysis of current trends, explores potential future opportunities, and includes predictive market forecasts. For more in-depth information and to access the full report, please visit -https://app.360iresearch.com/library/intelligence/brewers-spent-yeast